This page provides a step-by-step guide on the fastest route to getting started with BayesRTMB.
First, we will review the basic flow of rtmb_code() and
rtmb_model() using a simple binomial model. Next, we will
see the role of each block in a regression model. Finally, we will learn
how to use random effects and the Laplace approximation in a
hierarchical model.
Workflow Covered in this Page
The basic usage of BayesRTMB boils down to three main steps:
- Prepare the data
- Write the model using
rtmb_code() - Create the model object with
rtmb_model()and useoptimize(),sample(), orvariational()
Binomial Distribution Model
First, we will look at the basic flow of rtmb_code() and
rtmb_model() using the simplest binomial model. Here, we
deal with a situation where 6 successes were observed out of 10
trials.
library(BayesRTMB)
Trial <- 10
Y <- 6
data_list <- list(Trial = Trial, Y = Y)
model_code <- rtmb_code(
parameters={
theta=Dim(lower=0, upper=1)
},
model={
Y~binomial(Trial,theta)
theta ~ beta(1, 1)
}
)Creating the Model Object
rtmb_model() is a function that takes the data and model
definition and creates a model object for estimation.
The minimum required inputs are the data summarizing the
observations and the code created by
rtmb_code().
At this stage, estimation has not yet been performed. We will
subsequently run optimize(), sample(), and
variational() on the created model object.
mdl <- rtmb_model(data = data_list, code = model_code)## Pre-checking model code...
## Checking RTMB setup...MAP Estimation (Maximum A Posteriori)
optimize() performs MAP estimation. This is used when
you want to find the point estimate corresponding to
the mode of the posterior distribution. First, let’s run it without
additional options and check the output for estimates and interval
estimation.
fit_MAP <- mdl$optimize()
fit_MAP## Call:
## MAP Estimation via RTMB
##
## Negative Log-Posterior: 1.38
## Approx. Log Marginal Likelihood (Laplace): -0.90
##
## Point Estimates and 95% Wald CI:
## variable Estimate Std. Error Lower 95% Upper 95%
## theta 0.59976 0.15495 0.29720 0.84152 se_sampling option
If you specify se_sampling = TRUE, it calculates the
standard errors and 95% intervals via simulation using
the uncertainty of the estimation results. This method allows for
accurate 95% confidence intervals even for parameters with minimum or
maximum boundaries.
This is useful when you also want to see the confidence intervals for derived quantities.
fit_MAP <- mdl$optimize(se_sampling = TRUE)
fit_MAP## Call:
## MAP Estimation via RTMB
##
## Negative Log-Posterior: 1.38
## Approx. Log Marginal Likelihood (Laplace): -0.90
##
## Point Estimates and 95% Wald CI:
## variable Estimate Std. Error Lower 95% Upper 95%
## theta 0.59976 0.14563 0.29416 0.83985 Approximate Log Marginal Likelihood
In MAP estimation, an approximate log marginal likelihood
log_ml is calculated. This quantity is based on the
Laplace approximation and serves as a reference for
model comparison.
However, caution is needed if the posterior distribution is strongly skewed or multimodal.
fit_MAP$log_ml## [1] -2.328722MCMC Estimation
sample() performs MCMC sampling using NUTS. The default
values are sampling = 1000, warmup = 1000,
chains = 4, and thin = 1.
fit_mcmc <- mdl$sample(sampling = 1000,
warmup = 1000,
chains = 4,
thin = 1
)
fit_mcmc## variable mean sd map q2.5 q97.5 ess_bulk ess_tail rhat
## lp -3.33 0.74 -2.86 -5.43 -2.80 1712 1744 1.00
## theta 0.59 0.14 0.62 0.32 0.84 1609 1577 1.00 Saving and Loading MCMC Results (save_csv)
For models that take a long time to compute or when performing
large-scale sampling, it is useful to save the results to CSV files. By
specifying the save_csv argument during the execution of
sample() (or variational()), the sampling
results for each chain will be written to the specified directory.
# Save results to the "mcmc_out" directory with the name prefix "my_model"
fit_mcmc <- mdl$sample(
save_csv = list(name = "my_model", dir = "mcmc_out")
)The saved results can be read back into a model object later using
the read_mcmc_csv() function. This allows you to reuse
sampling results for plotting and summarization even after closing the
session or restarting R.
# Load saved files
# (mdl must be a model object created with the same settings)
fit_mcmc <- read_mcmc_csv(mdl, name = "my_model", dir = "mcmc_out")Visualizing Inference Results
To check the shape of the posterior distribution obtained via MCMC and its convergence status, BayesRTMB provides several visualization functions.
These functions can be used by passing the samples extracted with the
$draws() method from the model object.
# Extract posterior samples
samples <- fit_mcmc$draws("theta")
# Posterior density plot
plot_dens(samples)
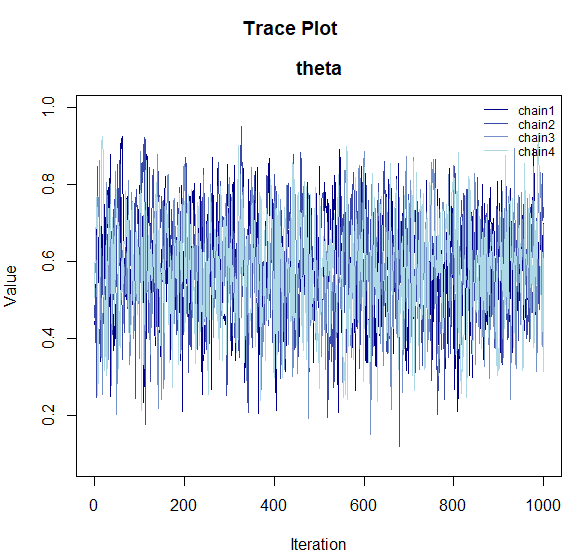
# Trace plot (checking convergence)
plot_trace(samples)
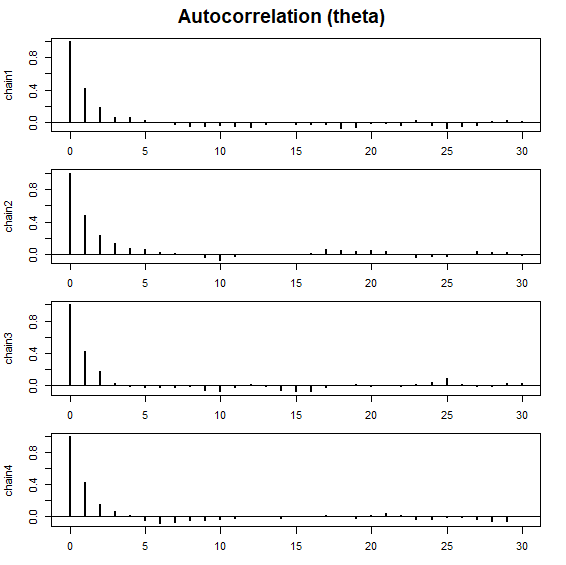
# Autocorrelation plot
plot_acf(samples)
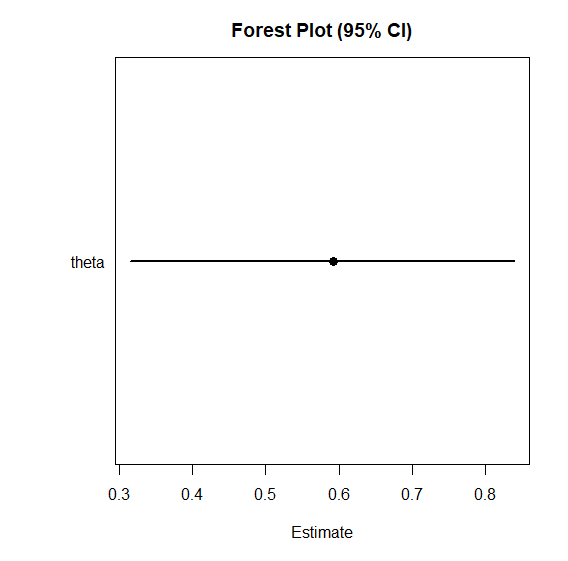
# Forest plot (overview of point estimates and intervals)
plot_forest(samples)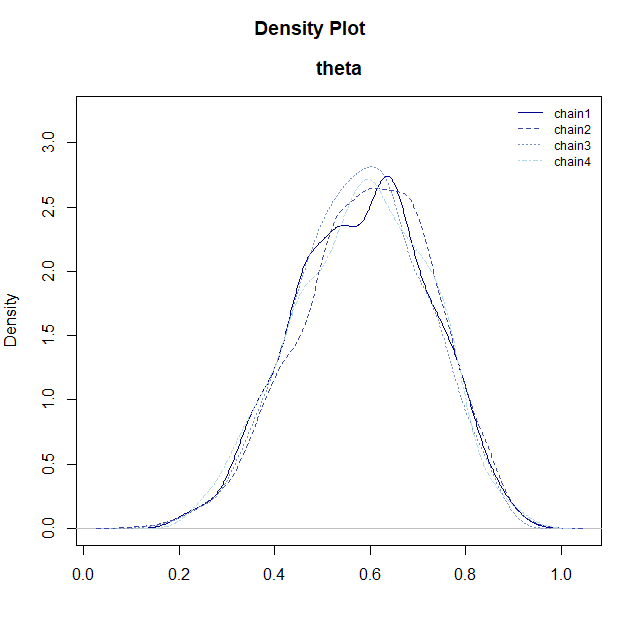
-
plot_dens(): Displays the posterior distribution density for each parameter. -
plot_trace(): Shows how each chain moved through the parameter space. If the chains are overlapping (resembling a “caterpillar”), convergence is likely. -
plot_acf(): Plots the autocorrelation of the MCMC samples. -
plot_forest(): Displays the point estimates (median) and 95% credible intervals for the parameters side-by-side.
Computing Log Marginal Likelihood via bridgesampling
By using bridgesampling(), you can estimate the
log marginal likelihood based on the MCMC samples. The
output also includes an estimation error, so if the error is large, you
can increase the number of samples to improve accuracy.
Once calculated, the result is saved in fit_mcmc$log_ml,
allowing it to be reused later on the same object.
fit_mcmc$bridgesampling()
fit_mcmc$log_ml## [1] -2.401682
## attr(,"error")
## [1] 0.001049984
## attr(,"ess")
## [1] 1204.976Regression Model
Next, let’s look at the role of each block in
rtmb_code() using a regression analysis as an example. In
this example, the flow is as follows:
-
setup: Organize data dimensions. -
parameters: Declare the parameters to be estimated. -
transform: Define the mean structuremu. -
model: Write the likelihood and prior distributions.
Note that for standard analyses such as regression or generalized
linear (mixed) models, you can use wrapper functions like
rtmb_lm or rtmb_glmer to automatically
generate and estimate models without writing code yourself. For details,
see Wrapper Functions.
Here, we will build a model that assigns normal distributions to the intercept and regression coefficients, and an exponential distribution to the residual standard deviation as priors. We will use the discussion data included in this package. This data contains the satisfaction of a discussion as the objective variable, and features like the amount of talk, discussion performance, conversation skills, and condition (discussion goal). Here, we will use the amount of talk, performance, and conversation skills.
data(discussion)
Y <- discussion$satisfaction
X_names <- c("talk","performance","skill")
X <- subset(discussion, select = X_names)
data_reg <- list(Y = Y, X = X)
code_reg <- rtmb_code(
setup = {
N <- length(Y)
P <- ncol(X)
},
parameters = {
alpha = Dim()
beta = Dim(P)
sigma = Dim(lower=0)
},
transform = {
mu = alpha + beta[1] * X[,1] + beta[2] * X[,2] + beta[3] * X[,3]
},
model = {
Y ~ normal(mu, sigma)
alpha ~ normal(0, 100)
beta ~ normal(0, 10)
sigma ~ exponential(1/10)
}
)Adding Initial Values to the Model
In rtmb_model(), you can provide initial values by
specifying init. Setting explicit initial values can often
help stabilize estimation in complex models. If you specify only some
parameters, the rest can be automatically filled in.
mdl_reg <- rtmb_model(data = data_reg,
code = code_reg,
init = list(alpha = 0, beta = c(0,0,0)))MCMC Estimation
For regression models as well, you can evaluate the entire posterior
distribution using sample().
mcmc_reg <- mdl_reg$sample()
mcmc_reg## variable mean sd map q2.5 q97.5 ess_bulk ess_tail rhat
## lp -411.42 1.70 -410.51 -415.77 -409.27 1099 1125 1.00
## alpha 1.51 0.24 1.52 1.03 1.98 805 842 1.01
## beta[1] 0.27 0.05 0.26 0.17 0.37 765 1223 1.00
## beta[2] 0.15 0.03 0.15 0.09 0.21 958 1107 1.01
## beta[3] 0.19 0.07 0.18 0.06 0.33 1004 1156 1.00
## sigma 0.90 0.04 0.90 0.83 0.98 984 1196 1.00
## mu[1] 2.96 0.10 2.96 2.76 3.16 1070 1095 1.00
## mu[2] 3.08 0.11 3.07 2.86 3.30 1254 2031 1.00
## mu[3] 2.96 0.10 2.96 2.76 3.16 1070 1095 1.00
## mu[4] 3.35 0.10 3.34 3.16 3.54 1489 2198 1.00 bayes_factor
By using bayes_factor(), you can compare the marginal
likelihood of the estimated model with a null model, calculating the
Bayes factor. For example, specifying
null_model = "beta[1]" internally creates and compares
against a null model where that coefficient is fixed at 0.
This is useful when you want to evaluate the presence or absence of an effect using a Bayesian approach.
bf_result <- mcmc_reg$bayes_factor(null_model = "beta[1]")
bf_result## Bayes Factor (BF12) : 1047.097
## Log Bayes Factor : 6.9538 (Approx. Error = 0.0051)
## Interpretation : Decisive evidence for Model 1 Hierarchical Linear Model
We will again use the discussion data for our
hierarchical model example. Here, we introduce random effects for each
group to represent individual differences.
In hierarchical models, centering explanatory variables and providing appropriate initial values can often be effective in improving the stability of the estimation.
data(discussion)
Y <- discussion$satisfaction
X_names <- c("talk","performance","skill")
X <- subset(discussion, select = X_names)
group <- discussion$group
data_hlm <- list(Y = Y, X = X, group = group)
code_hlm <- rtmb_code(
setup = {
N <- length(Y)
G <- length(unique(group))
P <- ncol(X)
},
parameters = {
alpha = Dim()
beta = Dim(P)
tau = Dim(lower=0)
sigma = Dim(lower=0)
r = Dim(G, random = TRUE)
},
transform = {
mu = alpha + X %*% beta + r[group] * tau
},
model = {
Y ~ normal(mu, sigma)
r ~ normal(0, 1)
alpha ~ normal(0, 100)
beta ~ normal(0, 10)
tau ~ exponential(1/10)
sigma ~ exponential(1/10)
}
)Model Setup
Specifying par_names and view when creating
the model makes the summary output easier to read.
par_names is used to assign display names to parameters,
and view is used to specify which variables should be
prioritized for display at the top in outputs like
summary().
In this example, we use par_names to name the regression
coefficients and view to ensure tau is
displayed above sigma. Setting these two options in advance
is convenient for improving output readability.
mdl_hlm <-
rtmb_model(
data = data_hlm,
code = code_hlm,
par_names = list(beta = X_names),
view = c("alpha", "beta", "tau", "sigma")
)MAP Estimation via Laplace Approximation
By setting optimize(laplace = TRUE) (which is the
default), you can estimate the fixed effects while marginalizing
(integrating out) the random effects using the Laplace
approximation. This often significantly reduces computation
time in hierarchical models and is useful for quickly grasping the
overall picture via MAP first.
opt_hlm <- mdl_hlm$optimize(laplace = TRUE)
opt_hlm## Call:
## MAP Estimation via RTMB
##
## Negative Log-Posterior: 399.48
## Approx. Log Marginal Likelihood (Laplace): -411.83
## Note: Random effects are stored in $random_effects
##
## Point Estimates and 95% Wald CI:
## variable Estimate Std. Error Lower 95% Upper 95%
## alpha 1.52450 0.25732 1.02015 2.02884
## beta[talk] 0.23487 0.05323 0.13054 0.33920
## beta[performance] 0.15451 0.03713 0.08175 0.22728
## beta[skill] 0.22613 0.05990 0.10874 0.34352
## tau 0.48512 0.06507 0.37298 0.63098
## sigma 0.74762 0.03752 0.67759 0.82488
## mu[1,1] 2.92366 0.34519 2.24711 3.60022
## mu[2,1] 3.14105 0.34856 2.45788 3.82422
## mu[3,1] 2.92366 0.34519 2.24711 3.60022
## mu[4,1] 3.09284 0.34706 2.41263 3.77306 Despite differences in priors, the Laplace approximation results are
almost identical to maximum likelihood estimation using
lme4. (Note: lme4 is not a dependency of
BayesRTMB. The following code is for comparison purposes
only and requires lme4 to be installed manually if you wish
to run it.)
library(lme4)
result <-
lmer(satisfaction ~ talk + performance + skill + (1|group),
data = discussion, REML = FALSE)
result |> summary()## Linear mixed model fit by maximum likelihood ['lmerMod']
## Formula: satisfaction ~ talk + performance + skill + (1 | group)
## Data: discussion
##
## AIC BIC logLik -2*log(L) df.resid
## 771.1 793.4 -379.6 759.1 294
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -3.15138 -0.51022 0.03797 0.54144 2.87725
##
## Random effects:
## Groups Name Variance Std.Dev.
## group (Intercept) 0.2357 0.4855
## Residual 0.5601 0.7484
## Number of obs: 300, groups: group, 100
##
## Fixed effects:
## Estimate Std. Error t value
## (Intercept) 1.52449 0.25735 5.924
## talk 0.23485 0.05286 4.443
## performance 0.15451 0.03714 4.160
## skill 0.22616 0.05960 3.795
##
## Correlation of Fixed Effects:
## (Intr) talk prfrmn
## talk -0.516
## performance -0.632 -0.056
## skill -0.387 -0.138 -0.020The estimation results for the random effects are saved separately in
random_effects.
opt_hlm$random_effects$Estimate |> hist()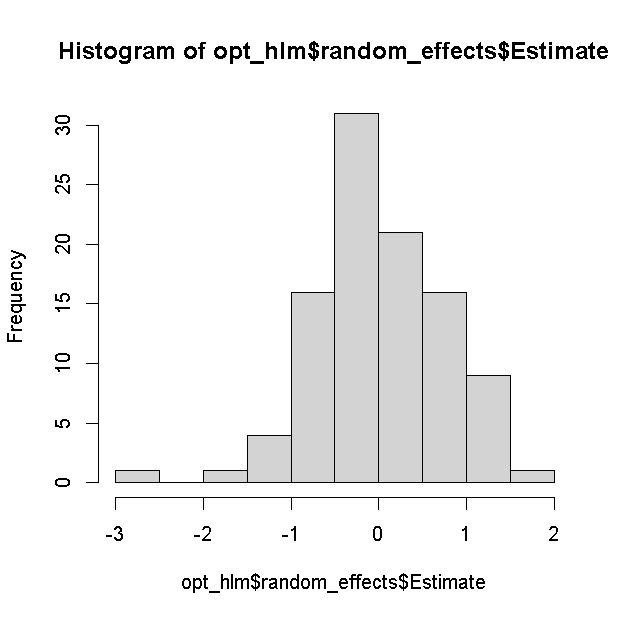
MCMC
Hierarchical models can also be estimated stably with MCMC via
sample(). If there are many chains or the model is heavy,
you can specify parallel = TRUE for parallel processing.
The first time you run parallel processing, setting up the workers takes
some time, but subsequent runs will launch faster (it is set to
FALSE by default).
You can increase warmup and sampling as
needed while checking convergence diagnostics.
mcmc_hlm <- mdl_hlm$sample(parallel = TRUE)
mcmc_hlm$summary()## variable mean sd map q2.5 q97.5 ess_bulk ess_tail rhat
## lp -503.48 11.43 -502.73 -527.43 -482.45 845 1883 1.00
## alpha 1.53 0.26 1.50 1.02 2.04 1885 2572 1.00
## beta[talk] 0.24 0.06 0.23 0.13 0.34 3292 3346 1.00
## beta[performance] 0.15 0.04 0.16 0.08 0.23 1724 2415 1.00
## beta[skill] 0.23 0.06 0.21 0.10 0.34 4128 3016 1.00
## tau 0.50 0.07 0.50 0.36 0.63 1532 1885 1.00
## sigma 0.76 0.04 0.76 0.69 0.84 2123 3211 1.00
## r[1] 0.02 0.66 0.00 -1.28 1.27 6016 2961 1.00
## r[2] -0.56 0.66 -0.47 -1.86 0.73 5403 2966 1.00
## r[3] -0.75 0.68 -0.77 -2.07 0.58 5041 3202 1.00 You can also run the Laplace approximation with MCMC. Although the estimation results do not change, it tends to reduce MCMC autocorrelation, making convergence more stable. However, the computation time increases, so it’s a trade-off. If MCMC converges sufficiently well, you might not need to explicitly use the Laplace approximation. However, it is sometimes convenient to do it automatically since comparing models with and without hierarchies using predictive metrics like WAIC can be difficult.
mcmc_hlm_l <- mdl_hlm$sample(laplace = TRUE, parallel = TRUE)ADVI
For complex hierarchical models, MAP might fail to find a solution,
and MCMC might take too long. Variational Bayes is useful when you want
to obtain an approximate solution in such cases. Variational Bayes can
also be parallelized. However, for complex models, judging convergence
can be difficult, so it’s important to check if it has converged using
the plot_elbo() method. Since Variational Bayes is
sensitive to initial values, providing rough initial values stabilizes
convergence.
You can also use the Laplace approximation in ADVI, though it slows
down the computation. method = "meanfield" assumes that the
posterior distribution components are all independent. Although the
credible intervals are estimated somewhat narrowly, it is perfectly
adequate for obtaining point estimates.
vb_hlm <- mdl_hlm$variational(
iter = 7000,
parallel = TRUE,
method = "meanfield",
laplace = TRUE
)
vb_hlm$summary(digits=5)## variable mean sd map q2.5 q97.5
## lp -403.59863 2.68487 -401.72373 -410.58114 -400.88726
## alpha 1.52112 0.06460 1.52966 1.39289 1.65062
## beta[talk] 0.23008 0.02011 0.22903 0.18939 0.26960
## beta[performance] 0.15445 0.01236 0.15815 0.12902 0.17917
## beta[skill] 0.22121 0.02962 0.23026 0.16046 0.27926
## tau 0.52022 0.06577 0.50972 0.40743 0.65876
## sigma 0.76221 0.03666 0.75816 0.69228 0.83982
## mu[1,1] 2.91530 0.04257 2.91892 2.83138 3.00157
## mu[2,1] 3.12764 0.06905 3.15095 2.99470 3.26365
## mu[3,1] 2.91530 0.04257 2.91892 2.83138 3.00157 Furthermore, if you set the method to fullrank, you can
estimate assuming a multivariate normal distribution where the posterior
distribution is not independent. Theoretically, specifying
laplace=TRUE and method = "fullrank" should
yield results similar to MAP.
vb_hlm <- mdl_hlm$variational(
iter = 7000,
parallel = TRUE,
method = "fullrank",
laplace = TRUE
)
vb_hlm$summary(digits=5)## variable mean sd map q2.5 q97.5
## lp -403.97866 2.20926 -402.90991 -409.23611 -401.18538
## alpha 1.53770 0.25051 1.56744 1.01054 2.02055
## beta[talk] 0.24155 0.05547 0.23247 0.13522 0.35168
## beta[performance] 0.15541 0.03070 0.14594 0.09867 0.21363
## beta[skill] 0.22281 0.06458 0.24644 0.09070 0.34811
## tau 0.51825 0.07037 0.52741 0.39687 0.66168
## sigma 0.75946 0.05066 0.76286 0.66550 0.85404
## mu[1,1] 2.94047 0.07042 2.92289 2.80572 3.08573
## mu[2,1] 3.14453 0.09346 3.14312 2.96019 3.32846
## mu[3,1] 2.94047 0.07042 2.92289 2.80572 3.08573 If some estimation results converge poorly, you can look only at the best ones. As long as the estimated ELBOs from the last 1000 iterations align horizontally in a random manner, it’s fine. If it’s sloping upwards, it might not have converged yet.
vb_hlm$plot_elbo(ests="best")
vb_hlm$summary()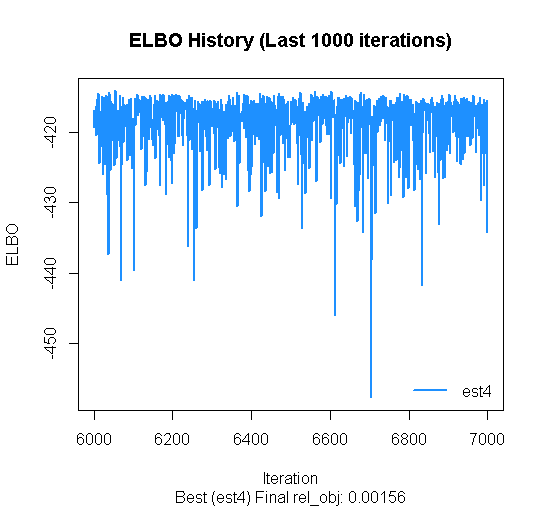
Model Comparison
Model comparison can be performed by calculating the log marginal likelihood.
Let’s compare the results of the regression analysis earlier with the hierarchical linear model. How much did the model improve by estimating the random effects?
Since MAP estimation can also calculate log marginal likelihoods, approximate model comparisons can be made.
opt_reg <- mdl_reg$optimize()
opt_reg$log_ml
opt_hlm$log_ml## > opt_reg$log_ml
## [1] -419.6358
## > opt_hlm$log_ml
## [1] -411.8348However, since the log marginal likelihood in MAP estimation is
strictly an approximation, it is better to calculate it using
bridgesampling on the MCMC estimation results for more
accuracy. In particular, because MAP uses the Laplace approximation when
there are random effects, its log marginal likelihood will differ
slightly from MCMC without Laplace approximation.
mcmc_reg$bridgesampling()
mcmc_hlm$bridgesampling()## > mcmc_reg$bridgesampling()
## Bridge Sampling Converged: LogML = -419.608 (Error = 0.0039, ESS = 830.6)
## [1] -419.6081
## attr(,"error")
## [1] 0.003854939
## attr(,"ess")
## [1] 830.623
## > mcmc_hlm$bridgesampling()
## Bridge Sampling Converged: LogML = -412.675 (Error = 0.0401, ESS = 760.2)
## [1] -412.6746
## attr(,"error")
## [1] 0.04009978
## attr(,"ess")
## [1] 760.1723MCMC with the Laplace approximation yields results closer to MAP.
mcmc_hlm_l$bridgesampling()## > mcmc_hlm_l$bridgesampling()
## Bridge Sampling Converged: LogML = -411.770 (Error = 0.0053, ESS = 994.9)
## [1] -411.7704
## attr(,"error")
## [1] 0.005318097
## attr(,"ess")
## [1] 994.8858GLMM
Generalized Linear Mixed Models (GLMMs) are also possible.
Since discussion satisfaction is measured on a 5-point scale, it can be treated as ordinal data. Therefore, let’s run a multilevel analysis assuming an ordered logistic distribution.
data(discussion)
Y <- discussion$satisfaction
X_names <- c("talk","performance","skill")
X <- subset(discussion, select = X_names)
group <- discussion$group
data_glmm <- list(Y = Y, X = X, group = group)
code_glmm <- rtmb_code(
setup = {
N <- length(Y)
G <- length(unique(group))
P <- ncol(X)
K <- length(unique(Y))
},
parameters = {
alpha = Dim(K-1, type = "ordered") # Ordered constrained vector
beta = Dim(P)
tau = Dim(lower=0)
r = Dim(G, random = TRUE)
},
transform = {
mu = X %*% beta + r[group] * tau
},
model = {
Y ~ ordered_logistic(mu, alpha) # Ordered logistic is available
r ~ normal(0, 1)
alpha ~ normal(0, 2.5)
beta ~ normal(0, 10)
tau ~ exponential(1/10)
}
)Loading the Model
mdl_glmm <- rtmb_model(data_glmm, code_glmm,
par_names = list(beta = X_names))GLMM via Laplace Approximation
Even for GLMMs, MAP estimation is possible by integrating out the random effects.
opt_glmm <- mdl_glmm$optimize()
opt_glmm## Call:
## MAP Estimation via RTMB
##
## Negative Log-Posterior: 391.24
## Approx. Log Marginal Likelihood (Laplace): -397.74
## Note: Random effects are stored in $random_effects
##
## Point Estimates and 95% Wald CI:
## variable Estimate Std. Error Lower 95% Upper 95%
## alpha[1] -0.30770 0.65457 -1.59063 0.97522
## alpha[2] 1.48733 0.60548 -0.32034 3.51175
## alpha[3] 4.25193 0.63446 1.98068 6.83335
## alpha[4] 6.50517 0.70195 3.81621 9.59934
## beta[talk] 0.50442 0.13449 0.24082 0.76802
## beta[performance] 0.32810 0.09502 0.14186 0.51434
## beta[skill] 0.49771 0.15633 0.19131 0.80411
## tau 1.27056 0.21778 0.90803 1.77784
## mu[1,1] 2.81917 0.93273 0.99105 4.64730
## mu[2,1] 3.31017 0.97502 1.39917 5.22117 Mixture Distribution Model
You can also estimate data that is a mixture of multiple distributions. First, let’s generate the data.
set.seed(123)
N <- 300 # Sample size
K <- 3 # Number of clusters
# True parameters
theta_true <- c(0.2, 0.5, 0.3) # Mixing ratio for each cluster (sum to 1)
mu_true <- c(-3, 0, 4) # Mean of each cluster
sigma_true <- c(0.5, 1.0, 0.8) # Standard deviation of each cluster
# Latent variable z (which cluster each data point comes from)
z <- sample(1:K, size = N, replace = TRUE, prob = theta_true)
# Observed data Y
Y <- numeric(N)
for (i in 1:N) {
Y[i] <- rnorm(1, mean = mu_true[z[i]], sd = sigma_true[z[i]])
}
Y |> hist()
data_mix <- list(Y = Y)Here is the code.
code_mix <- rtmb_code(
setup = {
K = 3 # Number of clusters
},
parameters = {
theta = Dim(K,type="simplex") # Positive vector summing to 1, common in mixing ratios
mu = Dim(K)
sigma = Dim(K, lower = 0)
},
model = {
mu ~ normal(0, 5)
sigma ~ exponential(1)
theta ~ dirichlet(rep(1, K)) # Dirichlet distribution as prior
Y ~ normal_mixture(theta, mu, sigma) # Built-in normal mixture distribution
}
)Model Preparation
mdl_mix <- rtmb_model(data_mix, code_mix)Setting Initial Values
Mixture distribution models often suffer from label switching if estimated directly. Label switching is a phenomenon where the cluster labeling changes across chains. While inevitable in MCMC, providing initial values makes it less likely to occur.
Therefore, we use MAP estimates. However, since mixture models are
highly dependent on initial values and often fall into local optima, we
run multiple MAP estimations. Setting num_estimate = 8 runs
it 8 times.
opt_mix <- mdl_mix$optimize(num_estimate = 8)## Starting optimization...
## Optimization run 8/8...
##
## Optimization Diagnostics per estimate:
## est1: Objective = 711.49, Code = 0 (Converged)
## est2: Objective = 711.49, Code = 0 (Converged)
## est3: Objective = 655.84, Code = 0 (Converged) <-- BEST
## est4: Objective = 695.90, Code = 0 (Converged)
## est5: Objective = 655.84, Code = 0 (Converged)
## est6: Objective = 675.00, Code = 0 (Converged)
## est7: Objective = 713.57, Code = 0 (Converged)
## est8: Objective = 676.45, Code = 0 (Converged)We run MCMC using this “Best” result as the initial value.
mcmc_mix <- mdl_mix$sample(parallel=TRUE, init = opt_mix$par)
mcmc_mix## variable mean sd map q2.5 q97.5 ess_bulk ess_tail rhat
## lp -664.57 2.05 -663.46 -669.30 -661.57 1717 2894 1.00
## theta[1] 0.29 0.03 0.29 0.24 0.35 3928 2829 1.00
## theta[2] 0.18 0.03 0.18 0.13 0.23 3274 2909 1.00
## theta[3] 0.53 0.04 0.53 0.46 0.60 2790 2986 1.00
## mu[1] 3.93 0.10 3.94 3.72 4.13 3551 2683 1.00
## mu[2] -2.98 0.09 -2.97 -3.15 -2.80 3210 3099 1.00
## mu[3] 0.03 0.11 0.02 -0.19 0.23 3506 3255 1.00
## sigma[1] 0.77 0.08 0.74 0.62 0.96 2724 2573 1.00
## sigma[2] 0.51 0.07 0.49 0.39 0.67 3459 3004 1.00
## sigma[3] 1.07 0.12 1.04 0.87 1.33 2758 2518 1.00 Summary
In this page, we learned the basic workflow of BayesRTMB through three types of models.
-
Simple Models: Understand the minimal setup using
rtmb_code()andrtmb_model(). -
Regression Models: Understand the roles of
setup,parameters,transform, andmodel. - Hierarchical Models: Understand the usage of random effects and Laplace approximation.
It is generally easier to first confirm the model’s behavior using MAP estimation, and then try MCMC or ADVI. For more details, viewing the Introduction along with the References will help you grasp the overall picture.